Baseline Correction in Python for Spectroscopy
Technique overview
Remove baseline drift from spectroscopy and chromatography traces before peak analysis or quantitative fitting.
Baseline drift can distort peak height, peak area, and fitted parameters in Raman, FTIR, UV-Vis, fluorescence, and chromatography data. A strong baseline can make a peak appear larger than it is or hide small features entirely. Baseline correction should be treated as a preprocessing model with assumptions, not as a cosmetic adjustment. This workflow compares polynomial fitting and asymmetric least squares so the corrected trace can be interpreted and reproduced.
Key points
- Remove baseline drift from spectroscopy and chromatography traces before peak analysis or quantitative fitting.
- Baseline drift can distort peak height, peak area, and fitted parameters in Raman, FTIR, UV-Vis, fluorescence, and chromatography data.
- A strong baseline can make a peak appear larger than it is or hide small features entirely.
- Baseline correction should be treated as a preprocessing model with assumptions, not as a cosmetic adjustment.
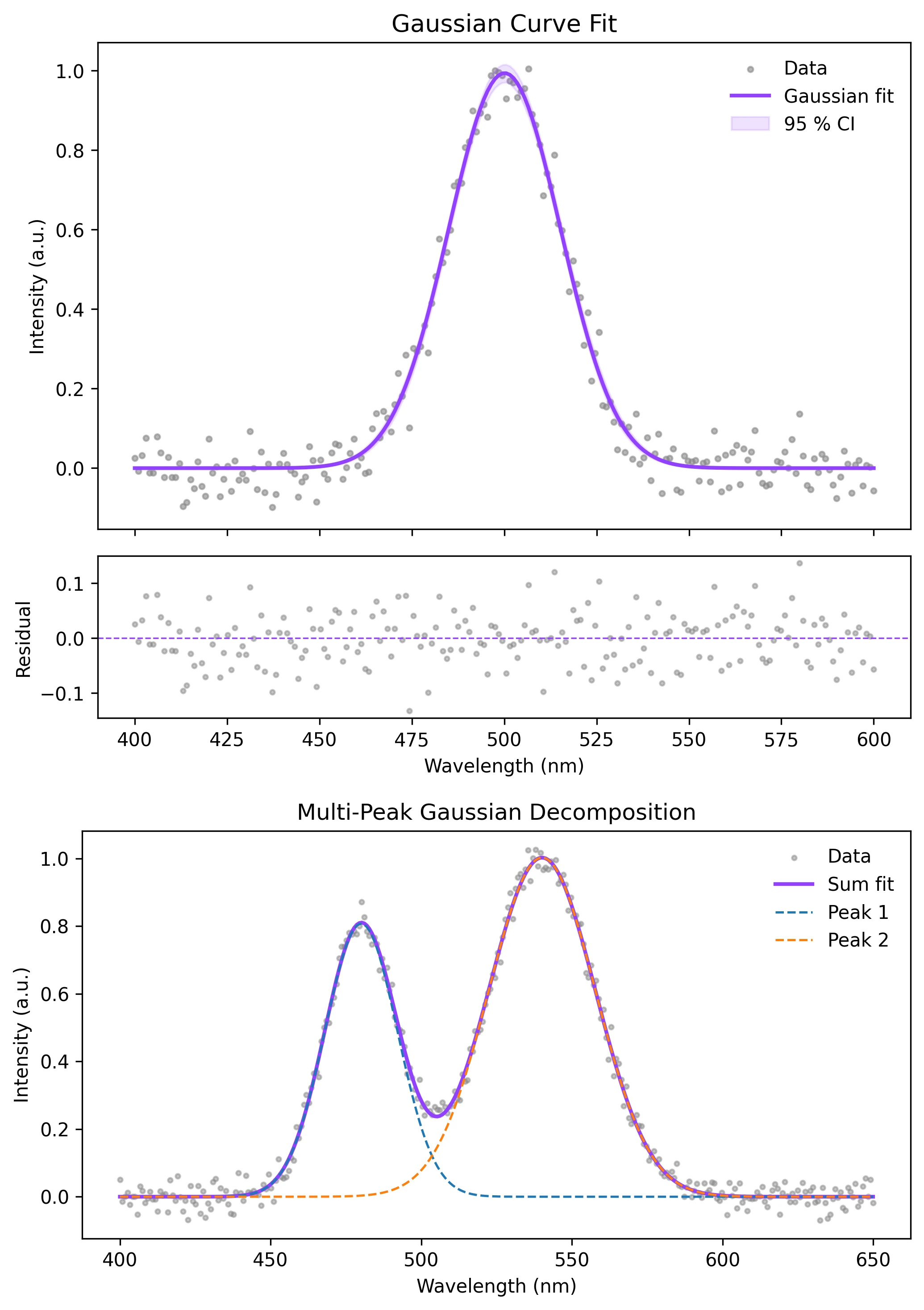
Example Visualization
Review the example first, then use the live editor below to run and customize the full workflow.
Mathematical Foundation
Baseline drift can distort peak height, peak area, and fitted parameters in Raman, FTIR, UV-Vis, fluorescence, and chromatography data.
Equation
corrected_signal = raw_signal - estimated_baselineParameter breakdown
When to use this technique
Use baseline correction before peak fitting, peak integration, or comparing spectra when background drift changes across samples.
Apply This Technique Now
Run this analysis workflow with AI in seconds. Use the prepared technique prompt or bring your own dataset.
View example prompt
"Correct the baseline of my spectroscopy data, show the raw trace versus corrected trace, and highlight the peaks after baseline removal"
How to apply this technique in 30 seconds
Generate
Run the example prompt and let AI generate this technique automatically.
Refine and Export
Adjust code or prompt, then export publication-ready figures.
Implementation Code
The core data processing logic. Copy this block and replace the sample data with your measurements.
import numpy as np
from scipy import sparse
from scipy.sparse.linalg import spsolve
def asymmetric_least_squares(y, lam=1e5, p=0.01, n_iter=10):
y = np.asarray(y, dtype=float)
length = len(y)
diff = sparse.diags([1, -2, 1], [0, 1, 2], shape=(length - 2, length))
weights = np.ones(length)
for _ in range(n_iter):
W = sparse.spdiags(weights, 0, length, length)
Z = W + lam * diff.T @ diff
baseline = spsolve(Z, weights * y)
weights = p * (y > baseline) + (1 - p) * (y < baseline)
return baseline
x = np.linspace(400, 1800, 800)
baseline_true = 0.0000004 * (x - 1100) ** 2 + 0.18
peaks = 0.8 * np.exp(-((x - 820) / 22) ** 2) + 0.55 * np.exp(-((x - 1320) / 35) ** 2)
raw = baseline_true + peaks + np.random.default_rng(5).normal(0, 0.025, size=x.size)
baseline = asymmetric_least_squares(raw, lam=1e6, p=0.01)
corrected = raw - baseline
print(f"Baseline range: {baseline.min():.3f} to {baseline.max():.3f}")Visualization Code
Complete matplotlib code for a publication-ready figure. Copy, paste into your notebook, and adjust labels to match your data.
import numpy as np
import matplotlib.pyplot as plt
from scipy import sparse
from scipy.sparse.linalg import spsolve
def asymmetric_least_squares(y, lam=1e6, p=0.01, n_iter=10):
y = np.asarray(y, dtype=float)
length = len(y)
D = sparse.diags([1, -2, 1], [0, 1, 2], shape=(length - 2, length))
weights = np.ones(length)
for _ in range(n_iter):
W = sparse.spdiags(weights, 0, length, length)
baseline = spsolve(W + lam * D.T @ D, weights * y)
weights = p * (y > baseline) + (1 - p) * (y < baseline)
return baseline
rng = np.random.default_rng(5)
x = np.linspace(400, 1800, 800)
baseline_true = 0.0000004 * (x - 1100) ** 2 + 0.18
peaks = 0.8 * np.exp(-((x - 820) / 22) ** 2) + 0.55 * np.exp(-((x - 1320) / 35) ** 2)
raw = baseline_true + peaks + rng.normal(0, 0.025, size=x.size)
baseline = asymmetric_least_squares(raw)
corrected = raw - baseline
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(8, 6), sharex=True)
ax1.plot(x, raw, color="#888888", lw=1, label="Raw spectrum")
ax1.plot(x, baseline, color="#9240ff", lw=2, label="Estimated baseline")
ax1.set_ylabel("Intensity")
ax1.set_title("Spectroscopy Baseline Correction")
ax1.legend(frameon=False)
ax2.plot(x, corrected, color="#111111", lw=1.1)
ax2.axhline(0, color="#9240ff", lw=1, ls=":")
ax2.set_xlabel("Wavenumber (cm^-1)")
ax2.set_ylabel("Corrected intensity")
ax1.spines[["top", "right"]].set_visible(False)
ax2.spines[["top", "right"]].set_visible(False)
plt.tight_layout()
plt.savefig("baseline_correction_spectroscopy.png", dpi=300, bbox_inches="tight")
plt.show()Polynomial Baseline for Peak-Free Regions
When you can identify baseline-only regions, a low-order polynomial fit is transparent and easy to report.
mask = ((x < 650) | ((x > 1000) & (x < 1150)) | (x > 1550)) coeff = np.polyfit(x[mask], raw[mask], deg=2) poly_baseline = np.polyval(coeff, x) poly_corrected = raw - poly_baseline
Common Errors and How to Fix Them
Baseline cuts through peaks
Why: The baseline model is too flexible or the asymmetry parameter is too high.
Fix: Increase lambda for a smoother baseline and reduce p so peaks are penalized less.
Negative corrected intensities are interpreted as real dips
Why: Baseline subtraction can shift noise below zero.
Fix: Inspect raw and corrected traces together and avoid over-interpreting small negative residuals.
Baseline parameters are not reported
Why: Preprocessing choices affect peak areas and fitted results.
Fix: Report the baseline method, lambda, p, polynomial degree, and excluded regions.
Frequently Asked Questions
Apply Baseline Correction in Python for Spectroscopy to Your Data
Upload your dataset and Plotivy generates the Python code, runs the analysis, and produces a publication-ready figure.
Generate Code for This TechniquePython Libraries
Quick Info
- Domain
- Signal Processing
- Typical Audience
- Spectroscopists and analytical chemists cleaning Raman, FTIR, UV-Vis, or chromatography data before peak quantification