Savitzky-Golay Smoothing in Python for Spectroscopy Data
Technique overview
Smooth spectroscopy and chromatography data while preserving peak shapes. Includes parameter selection guide for window length and polynomial order.
Noise is unavoidable in experimental science - a Raman spectrometer has shot noise, an FTIR instrument has detector noise, and an HPLC signal has electronic baseline drift. Savitzky-Golay filtering is the standard first-pass smoothing method because it preserves peak shapes, heights, and positions far better than a moving average. Instead of averaging neighbouring points equally, it fits a local polynomial to each window and uses the polynomial value at the center point, effectively acting as a low-pass filter that respects the underlying signal curvature. This page shows you how to choose the window length and polynomial order, visualize the effect of different parameters, and calculate smoothed derivatives for peak detection.
Key points
- Smooth spectroscopy and chromatography data while preserving peak shapes. Includes parameter selection guide for window length and polynomial order.
- Noise is unavoidable in experimental science - a Raman spectrometer has shot noise, an FTIR instrument has detector noise, and an HPLC signal has electronic baseline drift.
- Savitzky-Golay filtering is the standard first-pass smoothing method because it preserves peak shapes, heights, and positions far better than a moving average.
- Instead of averaging neighbouring points equally, it fits a local polynomial to each window and uses the polynomial value at the center point, effectively acting as a low-pass filter that respects the underlying signal curvature.
Example Visualization
Review the example first, then use the live editor below to run and customize the full workflow.
Mathematical Foundation
Noise is unavoidable in experimental science - a Raman spectrometer has shot noise, an FTIR instrument has detector noise, and an HPLC signal has electronic baseline drift.
Equation
y_smoothed[i] = sum_{j=-m}^{m} c_j * y[i+j]Parameter breakdown
When to use this technique
Use this when your raw signal is noisy but peak height, width, and position must stay physically meaningful. The implementation below uses window_length=15 and polyorder=3, which is a practical starting point for spectroscopy traces with moderate noise.
Apply This Technique Now
Run this analysis workflow with AI in seconds. Use the prepared technique prompt or bring your own dataset.
View example prompt
"Smooth my spectroscopy data using Savitzky-Golay filter, show raw vs smoothed overlay, and compare different window sizes to find the best setting"
How to apply this technique in 30 seconds
Generate
Run the example prompt and let AI generate this technique automatically.
Refine and Export
Adjust code or prompt, then export publication-ready figures.
Implementation Code
The core data processing logic. Copy this block and replace the sample data with your measurements.
import numpy as np
from scipy.signal import savgol_filter
# --- Simulated noisy spectrum (e.g., Raman) ---
np.random.seed(42)
x = np.linspace(200, 1800, 500)
# Two Lorentzian peaks on a baseline
peak1 = 0.8 / (1 + ((x - 520) / 15) ** 2)
peak2 = 0.5 / (1 + ((x - 1050) / 25) ** 2)
baseline = 0.1 + 0.0001 * (x - 1000)
y_clean = peak1 + peak2 + baseline
y_noisy = y_clean + np.random.normal(0, 0.03, size=x.shape)
# --- Apply Savitzky-Golay filter ---
window_length = 15 # must be odd
polyorder = 3 # polynomial degree
y_smooth = savgol_filter(y_noisy, window_length=window_length,
polyorder=polyorder)
# --- Quantify smoothing performance ---
rmse_noisy = np.sqrt(np.mean((y_noisy - y_clean) ** 2))
rmse_smooth = np.sqrt(np.mean((y_smooth - y_clean) ** 2))
print(f"RMSE before smoothing: {rmse_noisy:.5f}")
print(f"RMSE after smoothing: {rmse_smooth:.5f}")
print(f"Noise reduction: {(1 - rmse_smooth / rmse_noisy) * 100:.1f} %")Visualization Code
Complete matplotlib code for a publication-ready figure. Copy, paste into your notebook, and adjust labels to match your data.
import numpy as np
import matplotlib.pyplot as plt
from scipy.signal import savgol_filter
# --- Data ---
np.random.seed(42)
x = np.linspace(200, 1800, 500)
peak1 = 0.8 / (1 + ((x - 520) / 15) ** 2)
peak2 = 0.5 / (1 + ((x - 1050) / 25) ** 2)
y_clean = peak1 + peak2 + 0.1
y_noisy = y_clean + np.random.normal(0, 0.03, size=x.shape)
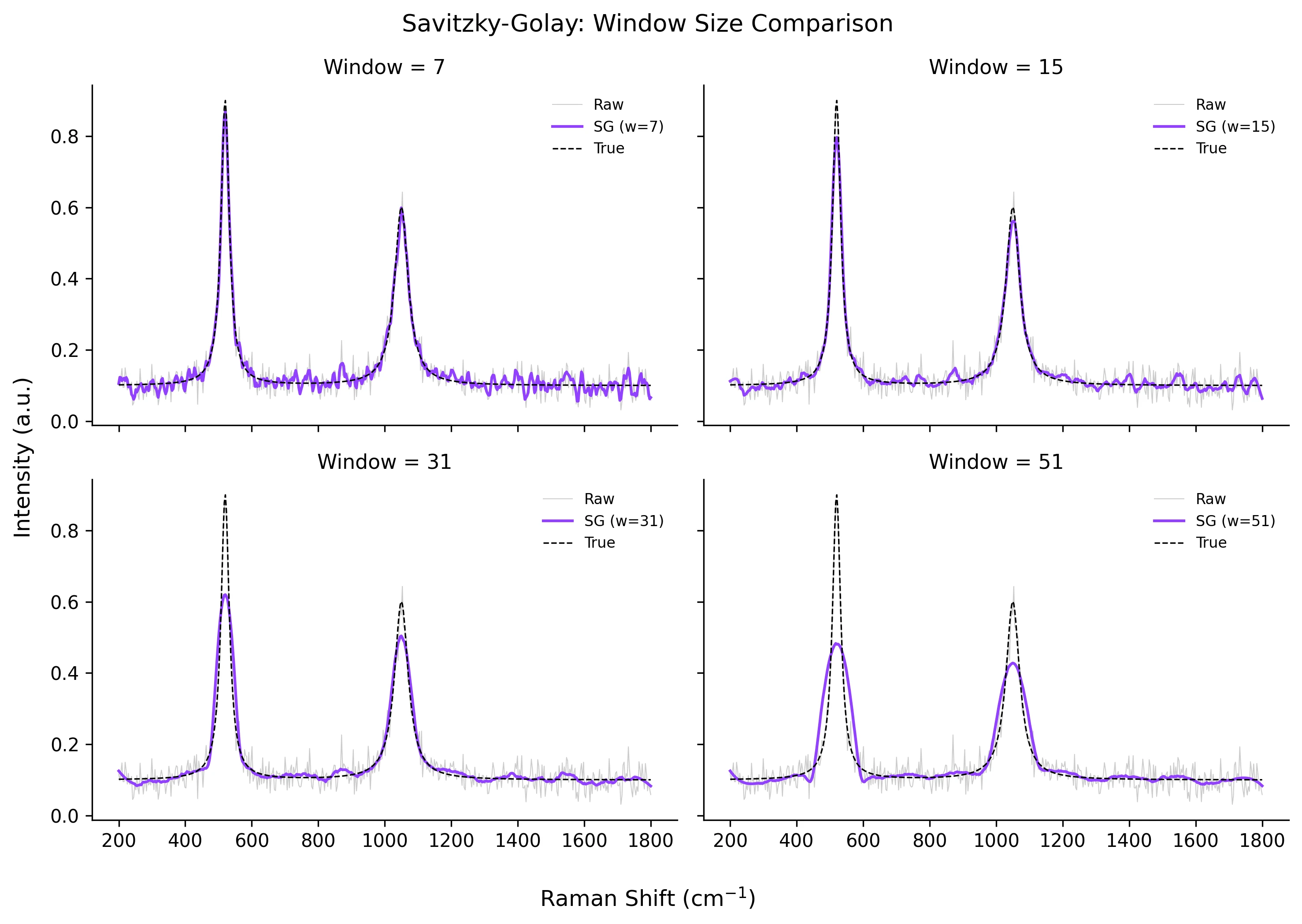
windows = [7, 15, 31, 51]
fig, axes = plt.subplots(2, 2, figsize=(10, 7), sharex=True, sharey=True)
for ax, w in zip(axes.flat, windows):
y_sm = savgol_filter(y_noisy, window_length=w, polyorder=3)
ax.plot(x, y_noisy, color='#cccccc', lw=0.5, label='Raw')
ax.plot(x, y_sm, color='#9240ff', lw=1.5, label=f'SG (w={w})')
ax.plot(x, y_clean, color='black', lw=0.8, ls='--', label='True')
ax.set_title(f'Window = {w}', fontsize=11)
ax.legend(frameon=False, fontsize=8)
ax.spines[['top', 'right']].set_visible(False)
fig.supxlabel('Raman Shift (cm$^{-1}$)')
fig.supylabel('Intensity (a.u.)')
fig.suptitle('Savitzky-Golay: Window Size Comparison', fontsize=13)
plt.tight_layout()
plt.savefig('savgol_comparison.png', dpi=300, bbox_inches='tight')
plt.show()Derivative Calculation with Savitzky-Golay
One of the most powerful features of Savitzky-Golay filtering is its ability to compute smooth derivatives simultaneously with the smoothing step. The first derivative highlights inflection points and helps resolve overlapping peaks. The second derivative is widely used in near-infrared spectroscopy to remove baseline offsets and enhance spectral features.
import numpy as np
import matplotlib.pyplot as plt
from scipy.signal import savgol_filter
np.random.seed(42)
x = np.linspace(200, 1800, 500)
peak = 0.8 / (1 + ((x - 520) / 15) ** 2) + 0.5 / (1 + ((x - 1050) / 25) ** 2)
y_noisy = peak + 0.1 + np.random.normal(0, 0.02, size=x.shape)
y_smooth = savgol_filter(y_noisy, 21, 3, deriv=0)
y_d1 = savgol_filter(y_noisy, 21, 3, deriv=1, delta=(x[1]-x[0]))
y_d2 = savgol_filter(y_noisy, 21, 3, deriv=2, delta=(x[1]-x[0]))
fig, axes = plt.subplots(3, 1, figsize=(8, 7), sharex=True)
axes[0].plot(x, y_noisy, color='#ccc', lw=0.6)
axes[0].plot(x, y_smooth, color='#9240ff', lw=1.5)
axes[0].set_ylabel('Intensity')
axes[0].set_title('Smoothed Spectrum', fontsize=11)
axes[1].plot(x, y_d1, color='#9240ff', lw=1.2)
axes[1].axhline(0, color='gray', lw=0.5, ls='--')
axes[1].set_ylabel('1st Derivative')
axes[1].set_title('First Derivative (peak positions at zero crossings)', fontsize=11)
axes[2].plot(x, y_d2, color='#9240ff', lw=1.2)
axes[2].axhline(0, color='gray', lw=0.5, ls='--')
axes[2].set_ylabel('2nd Derivative')
axes[2].set_title('Second Derivative (peaks become minima)', fontsize=11)
axes[2].set_xlabel('Raman Shift (cm$^{-1}$)')
plt.tight_layout()
plt.savefig('savgol_derivatives.png', dpi=300, bbox_inches='tight')
plt.show()Common Errors and How to Fix Them
ValueError: window_length must be odd
Why: scipy.signal.savgol_filter requires an odd window_length because the local polynomial is centered on the middle point of the window.
Fix: Always use odd values: 5, 7, 9, 11, etc. If you compute the window programmatically, use w = w if w % 2 == 1 else w + 1.
Window length is larger than the data array
Why: If your signal has fewer points than the window_length, the filter cannot construct any local fit.
Fix: Ensure window_length < len(data). For very short signals, reduce the window size or use a different smoothing method.
polyorder must be less than window_length
Why: A polynomial of degree d requires at least d+1 points. If polyorder >= window_length, the system is underdetermined.
Fix: Use polyorder = 2 or 3 for most applications. Only increase it if you have very wide windows and sharp features.
Edge artifacts (smoothed curve deviates near boundaries)
Why: At the edges, the filter has fewer neighbouring points and must extrapolate the polynomial, leading to artefacts.
Fix: Trim the first and last (window_length // 2) points from the smoothed result, or use mode="nearest" or mode="wrap" if appropriate for your data.
Peaks are broadened or reduced after smoothing
Why: A window that is too wide or a polynomial order that is too low effectively acts as a moving average, distorting narrow peaks.
Fix: Reduce the window_length until peak height and width match the original. Compare the smoothed curve to a reference or repeat measurements.
Frequently Asked Questions
Apply Savitzky-Golay Smoothing in Python for Spectroscopy Data to Your Data
Upload your dataset and Plotivy generates the Python code, runs the analysis, and produces a publication-ready figure.
Generate Code for This TechniquePython Libraries
Quick Info
- Domain
- Signal Processing
- Typical Audience
- Spectroscopists and analytical chemists processing Raman, FTIR, UV-Vis, or chromatography data who need noise reduction without peak distortion