Gaussian Curve Fitting in Python
Technique overview
Fit Gaussian peaks to spectroscopy, chromatography, or any bell-shaped data using scipy.optimize.curve_fit. Extract peak center, amplitude, and width with uncertainties.
Gaussian curve fitting is one of the most frequently encountered tasks in experimental science. Whether you are extracting emission line widths from a fluorescence spectrum, quantifying a chromatographic peak eluting from an HPLC column, or characterizing the point-spread function of an optical system, the underlying workflow is the same: define a Gaussian model, supply reasonable initial guesses, and let a nonlinear least-squares optimizer converge on the best-fit parameters. This page walks through the entire pipeline in Python using scipy.optimize.curve_fit, from raw data ingestion to publication-ready residual plots, so you can spend less time on boilerplate and more time interpreting your results.
Key points
- Fit Gaussian peaks to spectroscopy, chromatography, or any bell-shaped data using scipy.optimize.curve_fit. Extract peak center, amplitude, and width with uncertainties.
- Gaussian curve fitting is one of the most frequently encountered tasks in experimental science.
- Whether you are extracting emission line widths from a fluorescence spectrum, quantifying a chromatographic peak eluting from an HPLC column, or characterizing the point-spread function of an optical system, the underlying workflow is the same: define a Gaussian model, supply reasonable initial guesses, and let a nonlinear least-squares optimizer converge on the best-fit parameters.
- This page walks through the entire pipeline in Python using scipy.optimize.curve_fit, from raw data ingestion to publication-ready residual plots, so you can spend less time on boilerplate and more time interpreting your results.
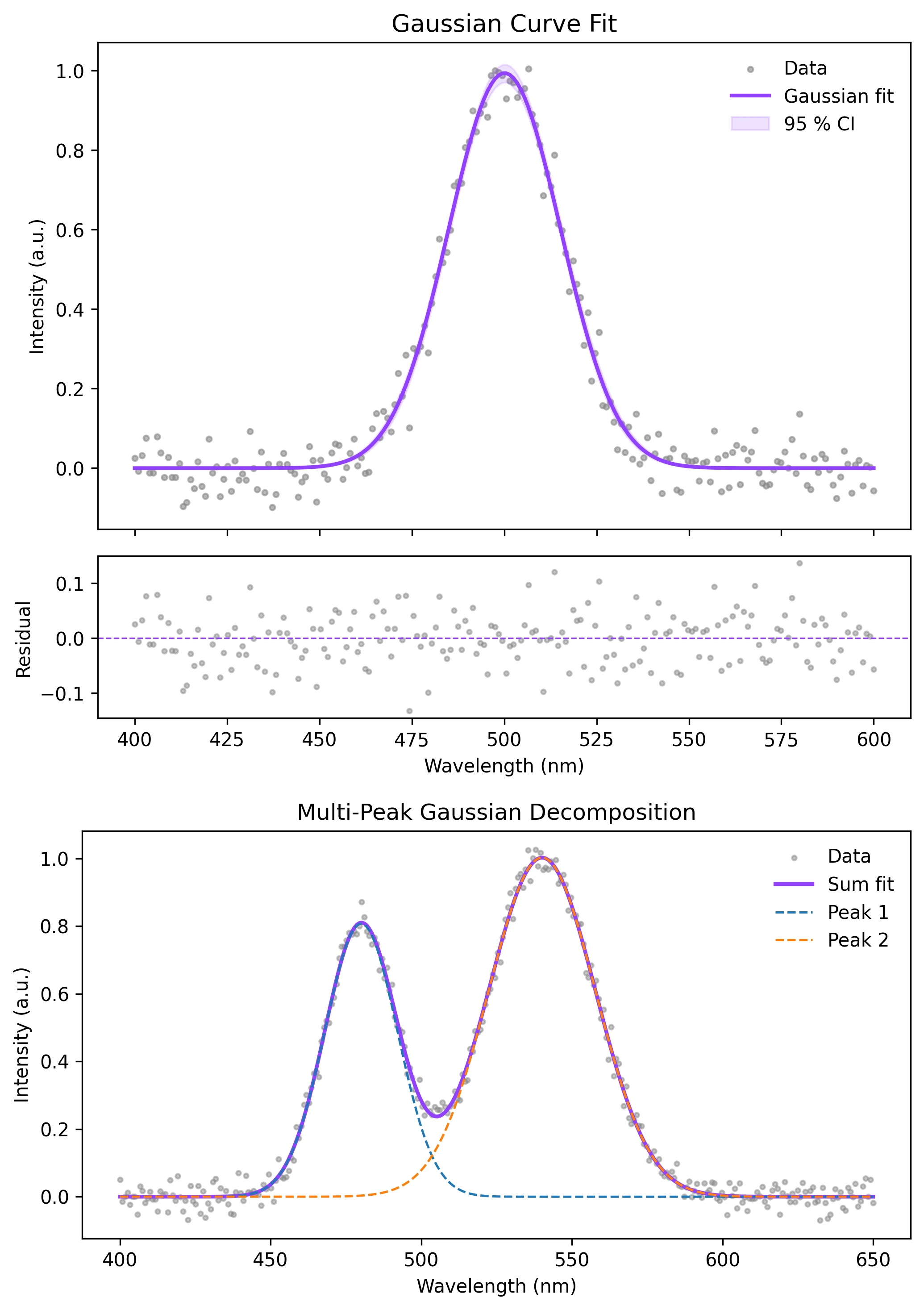
Example Visualization
Review the example first, then use the live editor below to run and customize the full workflow.
Mathematical Foundation
Gaussian curve fitting is one of the most frequently encountered tasks in experimental science.
Equation
f(x) = A * exp( -(x - mu)^2 / (2 * sigma^2) )Parameter breakdown
When to use this technique
Use a single Gaussian fit when your data contains one symmetric, bell-shaped peak riding on a flat or slowly varying baseline. If you see asymmetry or multiple overlapping peaks, consider a multi-peak model or a Voigt profile instead.
Apply This Technique Now
Run this analysis workflow with AI in seconds. Use the prepared technique prompt or bring your own dataset.
View example prompt
"Fit a Gaussian to my spectroscopy data and plot the result with residuals and parameter annotations in Nature style"
How to apply this technique in 30 seconds
Generate
Run the example prompt and let AI generate this technique automatically.
Refine and Export
Adjust code or prompt, then export publication-ready figures.
Implementation Code
The core data processing logic. Copy this block and replace the sample data with your measurements.
import numpy as np
from scipy.optimize import curve_fit
# --- Define the Gaussian model ---
def gaussian(x, amplitude, center, sigma):
return amplitude * np.exp(-(x - center) ** 2 / (2 * sigma ** 2))
# --- Simulated spectroscopy data (replace with your own) ---
np.random.seed(42)
x_data = np.linspace(400, 600, 200)
y_true = gaussian(x_data, amplitude=1.0, center=500, sigma=15)
y_data = y_true + np.random.normal(0, 0.05, size=x_data.shape)
# --- Initial parameter guesses ---
p0 = [y_data.max(), x_data[np.argmax(y_data)], 10.0]
# --- Fit ---
popt, pcov = curve_fit(gaussian, x_data, y_data, p0=p0)
perr = np.sqrt(np.diag(pcov))
print(f"Amplitude : {popt[0]:.4f} +/- {perr[0]:.4f}")
print(f"Center : {popt[1]:.4f} +/- {perr[1]:.4f}")
print(f"Sigma : {popt[2]:.4f} +/- {perr[2]:.4f}")
print(f"FWHM : {2.355 * popt[2]:.4f} +/- {2.355 * perr[2]:.4f}")Visualization Code
Complete matplotlib code for a publication-ready figure. Copy, paste into your notebook, and adjust labels to match your data.
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
def gaussian(x, amplitude, center, sigma):
return amplitude * np.exp(-(x - center) ** 2 / (2 * sigma ** 2))
# --- Data ---
np.random.seed(42)
x_data = np.linspace(400, 600, 200)
y_true = gaussian(x_data, 1.0, 500, 15)
y_data = y_true + np.random.normal(0, 0.05, size=x_data.shape)
popt, pcov = curve_fit(gaussian, x_data, y_data, p0=[1, 500, 10])
perr = np.sqrt(np.diag(pcov))
y_fit = gaussian(x_data, *popt)
residuals = y_data - y_fit
# --- 95 % confidence band ---
n_boot = 500
y_boot = np.zeros((n_boot, len(x_data)))
for i in range(n_boot):
p_sample = np.random.multivariate_normal(popt, pcov)
y_boot[i] = gaussian(x_data, *p_sample)
ci_low = np.percentile(y_boot, 2.5, axis=0)
ci_high = np.percentile(y_boot, 97.5, axis=0)
# --- Figure ---
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(7, 6),
gridspec_kw={'height_ratios': [3, 1]},
sharex=True)
ax1.scatter(x_data, y_data, s=8, color='#888888', alpha=0.6, label='Data')
ax1.plot(x_data, y_fit, color='#9240ff', linewidth=2, label='Gaussian fit')
ax1.fill_between(x_data, ci_low, ci_high, color='#9240ff', alpha=0.15,
label='95 % CI')
ax1.set_ylabel('Intensity (a.u.)')
ax1.legend(frameon=False)
ax1.set_title('Gaussian Curve Fit', fontsize=13)
ax2.scatter(x_data, residuals, s=6, color='#888888', alpha=0.5)
ax2.axhline(0, color='#9240ff', linewidth=0.8, linestyle='--')
ax2.set_xlabel('Wavelength (nm)')
ax2.set_ylabel('Residual')
plt.tight_layout()
plt.savefig('gaussian_fit.png', dpi=300, bbox_inches='tight')
plt.show()Multi-Peak Gaussian Fitting
When a spectrum contains two or three overlapping peaks, fit a sum of Gaussians simultaneously. The key challenge is providing sensible initial guesses for each component so the optimizer does not collapse multiple peaks into one.
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
def multi_gaussian(x, *params):
"""Sum of N Gaussians. params = [A1, mu1, sig1, A2, mu2, sig2, ...]"""
y = np.zeros_like(x)
for i in range(0, len(params), 3):
A, mu, sigma = params[i], params[i+1], params[i+2]
y += A * np.exp(-(x - mu) ** 2 / (2 * sigma ** 2))
return y
# --- Synthetic two-peak data ---
np.random.seed(7)
x = np.linspace(400, 650, 300)
y_true = (0.8 * np.exp(-(x - 480) ** 2 / (2 * 12 ** 2))
+ 1.0 * np.exp(-(x - 540) ** 2 / (2 * 18 ** 2)))
y = y_true + np.random.normal(0, 0.03, size=x.shape)
# --- Initial guesses for 2 peaks ---
p0 = [0.7, 475, 10, 0.9, 545, 15]
popt, pcov = curve_fit(multi_gaussian, x, y, p0=p0, maxfev=10000)
perr = np.sqrt(np.diag(pcov))
fig, ax = plt.subplots(figsize=(7, 4))
ax.scatter(x, y, s=6, color='#888', alpha=0.5, label='Data')
ax.plot(x, multi_gaussian(x, *popt), color='#9240ff', lw=2, label='Sum fit')
for i in range(0, len(popt), 3):
yi = popt[i] * np.exp(-(x - popt[i+1]) ** 2 / (2 * popt[i+2] ** 2))
ax.plot(x, yi, '--', lw=1.2, label=f'Peak {i//3 + 1}')
ax.legend(frameon=False)
ax.set_xlabel('Wavelength (nm)')
ax.set_ylabel('Intensity (a.u.)')
ax.set_title('Multi-Peak Gaussian Decomposition')
plt.tight_layout()
plt.savefig('multi_gaussian_fit.png', dpi=300, bbox_inches='tight')
plt.show()Common Errors and How to Fix Them
Fit returns the initial guesses unchanged
Why: Initial parameters are far from the true values, so the optimizer cannot descend into the correct minimum within the default iteration limit.
Fix: Estimate p0 from data: use max(y) for amplitude, x[argmax(y)] for center, and a rough FWHM/2.355 for sigma. Increase maxfev if needed.
pcov is inf or contains very large values
Why: The Jacobian is singular at the solution, often because the model is overparameterised or the data range does not constrain all parameters.
Fix: Reduce the number of free parameters, add bounds (e.g., sigma > 0), or collect more data points across the peak.
TypeError: only arrays of size 1 can be converted to Python scalars
Why: x_data or y_data were passed as lists of lists, DataFrames, or had mismatched shapes.
Fix: Convert inputs to 1-D NumPy arrays with np.asarray(x).flatten() before calling curve_fit.
Fitted sigma is unreasonably large (overfitting the baseline)
Why: The optimizer is pushing sigma to extremely large values to approximate a constant background offset.
Fix: Subtract or model the baseline before fitting, or add a constant offset parameter to the model: f(x) = A*exp(...) + c.
Negative amplitude in the fit result
Why: Without parameter bounds, curve_fit may flip the sign of A if the initial guess sign does not match the data.
Fix: Set bounds=([0, -np.inf, 0], [np.inf, np.inf, np.inf]) to constrain A and sigma to positive values.
Frequently Asked Questions
Learn More Before You Run It
Apply Gaussian Curve Fitting in Python to Your Data
Upload your dataset and Plotivy generates the Python code, runs the analysis, and produces a publication-ready figure.
Generate Code for This TechniquePython Libraries
Quick Info
- Domain
- Curve Fitting
- Typical Audience
- Spectroscopists, analytical chemists, and physicists fitting bell-shaped peaks to experimental data