4-Parameter Logistic Curve Fitting in Python
Technique overview
Fit 4-parameter logistic curves for assay standard curves, ELISA data, and dose-response experiments with EC50 or IC50 reporting.
The 4-parameter logistic model is the workhorse for ELISA standard curves, receptor binding assays, qPCR calibration, and concentration-response experiments. It captures the lower plateau, upper plateau, slope, and midpoint of a sigmoidal curve, which makes it more flexible than a straight line while still remaining interpretable. The practical challenge is fitting the model stably: concentrations usually need a log-scaled x-axis, initial guesses must be sensible, and residuals should be checked because a visually smooth sigmoid can still hide systematic assay bias.
Key points
- Fit 4-parameter logistic curves for assay standard curves, ELISA data, and dose-response experiments with EC50 or IC50 reporting.
- The 4-parameter logistic model is the workhorse for ELISA standard curves, receptor binding assays, qPCR calibration, and concentration-response experiments.
- It captures the lower plateau, upper plateau, slope, and midpoint of a sigmoidal curve, which makes it more flexible than a straight line while still remaining interpretable.
- The practical challenge is fitting the model stably: concentrations usually need a log-scaled x-axis, initial guesses must be sensible, and residuals should be checked because a visually smooth sigmoid can still hide systematic assay bias.
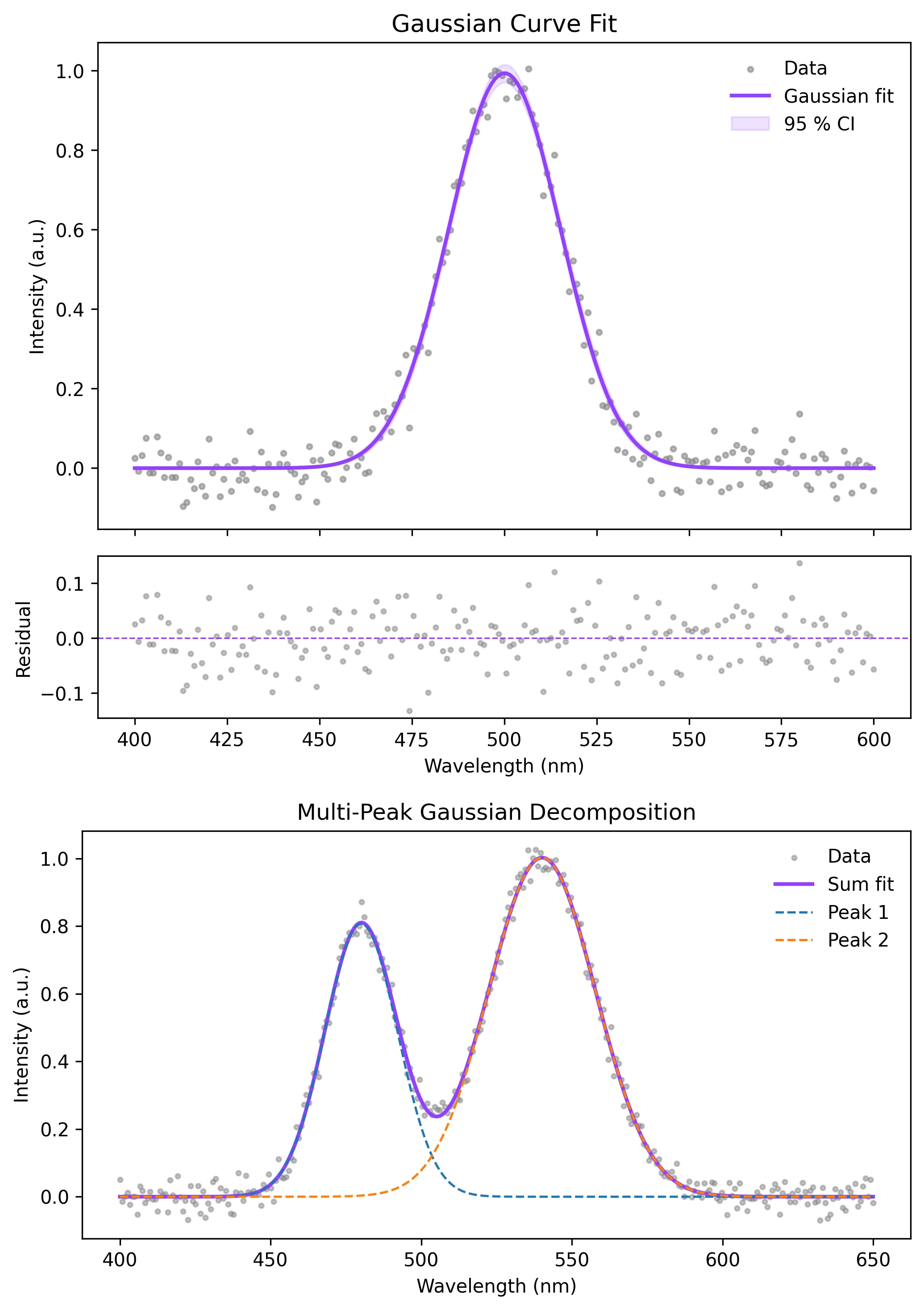
Example Visualization
Review the example first, then use the live editor below to run and customize the full workflow.
Mathematical Foundation
The 4-parameter logistic model is the workhorse for ELISA standard curves, receptor binding assays, qPCR calibration, and concentration-response experiments.
Equation
y = bottom + (top - bottom) / (1 + (x / EC50)^hill_slope)Parameter breakdown
When to use this technique
Use a 4PL model for monotonic sigmoidal assay curves with clear lower and upper plateaus. If one plateau is not observed, constrain that parameter from controls or report the estimate cautiously.
Apply This Technique Now
Run this analysis workflow with AI in seconds. Use the prepared technique prompt or bring your own dataset.
View example prompt
"Fit a 4-parameter logistic model to my assay data, report the EC50 with confidence intervals, show raw points with the fitted curve, and include residual diagnostics"
How to apply this technique in 30 seconds
Generate
Run the example prompt and let AI generate this technique automatically.
Refine and Export
Adjust code or prompt, then export publication-ready figures.
Implementation Code
The core data processing logic. Copy this block and replace the sample data with your measurements.
import numpy as np
from scipy.optimize import curve_fit
def four_pl(x, bottom, top, ec50, hill_slope):
return bottom + (top - bottom) / (1 + (x / ec50) ** hill_slope)
np.random.seed(42)
concentration = np.logspace(-2, 2, 16)
response_true = four_pl(concentration, bottom=0.08, top=1.05, ec50=3.2, hill_slope=-1.4)
response = response_true + np.random.normal(0, 0.04, size=concentration.size)
p0 = [response.min(), response.max(), np.median(concentration), -1.0]
bounds = ([0, 0, concentration.min() / 10, -5], [2, 2, concentration.max() * 10, 5])
popt, pcov = curve_fit(four_pl, concentration, response, p0=p0, bounds=bounds, maxfev=10000)
perr = np.sqrt(np.diag(pcov))
for name, value, err in zip(["bottom", "top", "EC50", "hill_slope"], popt, perr):
print(f"{name:10s}: {value:.4g} +/- {err:.2g}")Visualization Code
Complete matplotlib code for a publication-ready figure. Copy, paste into your notebook, and adjust labels to match your data.
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
def four_pl(x, bottom, top, ec50, hill_slope):
return bottom + (top - bottom) / (1 + (x / ec50) ** hill_slope)
np.random.seed(42)
x = np.logspace(-2, 2, 16)
y = four_pl(x, 0.08, 1.05, 3.2, -1.4) + np.random.normal(0, 0.04, size=x.size)
p0 = [y.min(), y.max(), np.median(x), -1.0]
popt, pcov = curve_fit(four_pl, x, y, p0=p0, bounds=([0, 0, 1e-4, -5], [2, 2, 1e4, 5]))
x_fit = np.logspace(np.log10(x.min()), np.log10(x.max()), 300)
y_fit = four_pl(x_fit, *popt)
residuals = y - four_pl(x, *popt)
fig, (ax, ax_res) = plt.subplots(2, 1, figsize=(7, 6), sharex=True,
gridspec_kw={"height_ratios": [3, 1]})
ax.scatter(x, y, color="#111111", s=32, label="Data")
ax.plot(x_fit, y_fit, color="#9240ff", lw=2.2, label="4PL fit")
ax.axvline(popt[2], color="#9240ff", ls=":", lw=1.2, label=f"EC50 = {popt[2]:.2g}")
ax.set_xscale("log")
ax.set_ylabel("Response")
ax.set_title("4-Parameter Logistic Curve Fit")
ax.legend(frameon=False)
ax_res.axhline(0, color="#888888", lw=1)
ax_res.scatter(x, residuals, color="#111111", s=24)
ax_res.set_xscale("log")
ax_res.set_xlabel("Concentration")
ax_res.set_ylabel("Residual")
plt.tight_layout()
plt.savefig("four_parameter_logistic_fit.png", dpi=300, bbox_inches="tight")
plt.show()Weighted 4PL for Heteroscedastic Assay Data
Assay variance often changes across concentration. If high-response wells are more variable than low-response wells, pass sigma values to curve_fit and set absolute_sigma=True so the fit reflects replicate uncertainty rather than treating every point equally.
# Example: replicate standard deviations from technical replicates
sigma = np.maximum(0.03, 0.08 * response)
popt_w, pcov_w = curve_fit(
four_pl,
concentration,
response,
p0=p0,
sigma=sigma,
absolute_sigma=True,
bounds=bounds,
maxfev=10000,
)
print(f"Weighted EC50: {popt_w[2]:.4g}")Common Errors and How to Fix Them
EC50 estimate is outside the tested concentration range
Why: The curve does not include enough of the transition region or one plateau is missing.
Fix: Extend the dilution series around the apparent midpoint or constrain the missing plateau from controls.
Fit flips upward when the assay should decrease
Why: The Hill slope sign is unconstrained and the initial guess points the optimizer toward the opposite curve direction.
Fix: Use a negative initial Hill slope for inhibitory curves and set bounds that match the expected direction.
Residuals curve systematically across the dilution series
Why: The 4PL model may be too simple or the data includes matrix effects, hook effect, or saturation artifacts.
Fix: Inspect controls, remove invalid points only with documented justification, or compare against a 5PL model.
Frequently Asked Questions
Python Statistics Code Templates for Biologists
Get 10 complete, runnable Python code snippets for the most common biological statistical tests, each with explanations and publication-ready figure output. Stop fighting with matplotlib!
Python Libraries
Quick Info
- Domain
- Biological
- Typical Audience
- Biologists, pharmacologists, and assay scientists quantifying standard curves, binding assays, or concentration-response relationships