PhD Figure Hell: How to Make Publication-Ready Figures Without Losing Your Mind (or Your Weekend)

It is Friday at 4:47 PM. Your supervisor sends you a message: "Can you finalize the figures this weekend? We are submitting Monday."
You think: no problem. The data is done. The analysis is done. How hard can a few figures be?
Seventy-two hours later you are awake at 3 AM, staring at a matplotlib window, wondering why the axis labels vanish when you export to PDF. The journal wants 300 DPI TIFF files with embedded fonts. Your legend covers the only interesting data point. Reviewer 2 will definitely notice the colors are indistinguishable in grayscale. And somewhere, deep in your heart, you know there are no error bars on Figure 4.
Every researcher has lived some version of this weekend. Figure formatting is not technically difficult - it is tedious, poorly documented, and unreasonably time-consuming. The gap between "I have a working plot in my Jupyter notebook" and "this is accepted by Nature" is about forty Stack Overflow tabs wide.
This guide is for you. We will walk through the five figure disasters that account for roughly 90% of formatting pain, give you the exact code to fix each one, and then talk honestly about how to build a workflow that does not steal your weekends.
What You'll Learn
1.The Five Figure Disasters (and How to Fix Them)
2.The Figure Workflow That Actually Works
3.Let Us Talk About Time
4.The Honest Recommendation
5.Your Figures Should Not Be the Hard Part
The Five Figure Disasters (and How to Fix Them)
These are not edge cases. These are the reasons journals send back revision requests before anyone even reads your results section. Each one is easy to fix once you know where to look.
Disaster 1: Your Figures Are 72 DPI (and the Journal Wants 300)
This is the single most common reason figures get rejected before peer review even begins. Matplotlib's default screen resolution is 100 DPI. If you use plt.savefig("fig.png") without specifying DPI, the output looks fine on your monitor but prints like a photograph taken through a wet window. Most journals - including Nature, Science, and every Elsevier title - require a minimum of 300 DPI for raster images.
The fix is one argument:
Updating preview...
Execution Error
Edit the code, then press Run Code to refresh the preview
Preparing preview
Running once automatically on first load
Learn by Experimenting
This is a safe playground for learning! Try changing:
- • Colors: Modify color values to see different palettes
- • Numbers: Adjust sizes, positions, or data ranges
- • Labels: Update titles, axis names, or legends
Edit the code, run it, then open the full data visualization tool to continue with your own dataset.
Always export both a raster (TIFF or PNG at 300+ DPI) and a vector (PDF or SVG) version. The vector is your insurance policy - it can be rescaled to any size without quality loss. Check your target journal's figure requirements before you export.
Disaster 2: Your Fonts Disappear at Print Size
Matplotlib's default font size is 10 points. That sounds reasonable until your figure gets shrunk to a single column width (roughly 3.5 inches) in the published PDF. Suddenly your axis labels are 6-point text - smaller than the fine print on a credit card agreement. Tick labels become illegible. The title might as well not exist.
Set your defaults once at the top of every script:
Updating preview...
Execution Error
Edit the code, then press Run Code to refresh the preview
Preparing preview
Running once automatically on first load
Learn by Experimenting
This is a safe playground for learning! Try changing:
- • Colors: Modify color values to see different palettes
- • Numbers: Adjust sizes, positions, or data ranges
- • Labels: Update titles, axis names, or legends
Edit the code, run it, then open the full data visualization tool to continue with your own dataset.
The rule of thumb: if your figure will be printed at single-column width, your smallest text should be at least 8 points in the final output. Start at 12-14 in your script and adjust from there. Arial and Helvetica are the safest choices - practically every journal accepts them, and they embed cleanly in PDFs.
Disaster 3: Your Colors Look Identical in Grayscale
About 8% of men and 0.5% of women have some form of color vision deficiency. Many journals still print in grayscale or charge extra for color pages. If your red and green lines become the same shade of gray, you have lost a significant chunk of your audience - including, potentially, a reviewer.
Use a colorblind-safe palette and vary line styles:
Updating preview...
Execution Error
Edit the code, then press Run Code to refresh the preview
Preparing preview
Running once automatically on first load
Learn by Experimenting
This is a safe playground for learning! Try changing:
- • Colors: Modify color values to see different palettes
- • Numbers: Adjust sizes, positions, or data ranges
- • Labels: Update titles, axis names, or legends
Edit the code, run it, then open the full data visualization tool to continue with your own dataset.
This palette comes from Bang Wong's 2011 paper in Nature Methods and is designed to remain distinguishable across all common forms of color blindness. Combine it with distinct line styles or markers for a figure that works in color, in grayscale, and on every screen. For more on choosing the right scatter plot or line chart visual encoding, see our technique guides.
Disaster 4: Your Legend Is Covering the Data
Matplotlib's loc="best" is a well-meaning lie. It picks the spot with the least overlap, but on a busy plot "least overlap" still means "covering something important." The classic fix is to move the legend outside the axes entirely. It uses a bit more whitespace, but nobody will ever email you to say "I cannot see the legend."
Place the legend outside the plot area:
Updating preview...
Execution Error
Edit the code, then press Run Code to refresh the preview
Preparing preview
Running once automatically on first load
Learn by Experimenting
This is a safe playground for learning! Try changing:
- • Colors: Modify color values to see different palettes
- • Numbers: Adjust sizes, positions, or data ranges
- • Labels: Update titles, axis names, or legends
Edit the code, run it, then open the full data visualization tool to continue with your own dataset.
The bbox_to_anchor tuple positions the legend relative to the axes. (1.02, 1) means "just past the right edge, aligned to the top." Combined with bbox_inches="tight", this ensures the legend is included in the exported file without clipping. For figures with many categories, consider placing the legend below the plot: bbox_to_anchor=(0.5, -0.15), loc="upper center".
Disaster 5: There Are No Error Bars (and Reviewer 2 Noticed)
Nothing invites a "where did you get that number?" comment faster than a bar chart with no error bars. Whether you report standard error of the mean (SEM) or standard deviation (SD) depends on your field and your question - SEM conveys precision of the mean estimate, SD conveys variability of individual measurements. Either way, you must show something.
Add error bars properly:
Updating preview...
Execution Error
Edit the code, then press Run Code to refresh the preview
Preparing preview
Running once automatically on first load
Learn by Experimenting
This is a safe playground for learning! Try changing:
- • Colors: Modify color values to see different palettes
- • Numbers: Adjust sizes, positions, or data ranges
- • Labels: Update titles, axis names, or legends
Edit the code, run it, then open the full data visualization tool to continue with your own dataset.
Always state in your figure caption whether error bars represent SEM, SD, or confidence intervals, and specify n (sample size). A bar chart without error bars is an opinion, not a result. For small sample sizes (n < 10), consider overlaying individual data points as a scatter plot - it is more transparent and increasingly expected by reviewers.
The Figure Workflow That Actually Works
The five fixes above solve individual problems. But the real time-saver is building a workflow that prevents those problems from appearing in the first place. After watching hundreds of researchers go through this process, here is what works:
Step 1: Sketch on Paper First
Before you open Python, grab a piece of paper and draw what you want the figure to communicate. Not how it should look - what story it tells. Which comparison matters? Where should the reader's eye go first? Ten minutes of sketching saves two hours of rearranging subplots.
Step 2: Pick the Right Tool for the Job
Try it
Try it now: review your figure before submission
Upload your current plot and get an AI critique with concrete fixes for clarity, typography, color, and journal readiness.
Open AI Figure Reviewer →Newsletter
Get a weekly Python plotting tip
One concise tip each week for cleaner, faster scientific figures. Built for researchers who publish.
Matplotlib is powerful but verbose. Plotly is interactive but can be overkill for static journal figures. GraphPad Prism is familiar to biologists but limited in customization. The best tool is the one where you spend time thinking about your data instead of fighting the API. We are biased, obviously, but Plotivy was built specifically for this gap: researchers who know what they want but do not want to become Python developers to get it.
Step 3: Code It Once, Never Touch It Again
The single biggest mistake is making figures interactively - dragging elements around in a GUI, tweaking colors by eye, manually adjusting axis ranges. The moment your data changes (and it will change), you have to redo everything from scratch. Write a script. Save the script next to the data. When Reviewer 2 asks you to add a panel, you re-run the script with one modification instead of recreating the figure from memory. This is the core principle behind reproducible research figures.
Step 4: Export at Final Journal Specs from Day One
Do not wait until submission day to figure out the journal's requirements. Set your figsize, dpi, and font sizes at the start of the project according to your target journal. Single-column width is typically 3.3-3.5 inches, double-column is 6.5-7 inches. Bake these into a template script and reuse it across every figure in the paper.
Let Us Talk About Time
Here is arithmetic that nobody does until it is too late.
Figure Time Budget Across a Typical PhD
- Raw matplotlib, manual formatting: 4-8 hours per complex figure (multi-panel, custom styling, journal-specific export)
- With a template library (e.g., seaborn defaults): 2-4 hours per figure
- With Plotivy: ~15 minutes per figure (describe what you want, adjust, export)
A typical PhD thesis and its associated papers produce roughly 30-50 publication-quality figures. Let us be conservative and say 30.
- Manual approach: 30 figures x 4-8 hours = 120-240 hours (that is 5-10 full weeks of work, spread across your entire PhD)
- With Plotivy: 30 figures x 0.25 hours = 7.5 hours
The difference is roughly 112-232 hours. That is an entire paper. Or a holiday. Or both.
These numbers are not marketing fiction. They are based on what we hear from researchers every week. The formatting part - DPI, fonts, colors, legends, spacing - is where time vanishes. The scientific thinking part (choosing the right plot, deciding what to emphasize) takes the same amount of time regardless of the tool.
The Honest Recommendation
We built Plotivy, so take everything below with appropriate skepticism. But here is a genuinely honest assessment of when to use what.
Use GraphPad Prism if you are in biology or clinical research, your lab already has a license, and your figures are standard plot types (bar charts, survival curves, dose-response). Prism is excellent for what it does. Its limitations show up when you need unusual customization or when you want to automate figure generation across many datasets.
Use R with ggplot2 if you are in ecology, epidemiology, or any field where R is the standard. The grammar of graphics is beautiful once you learn it, and the ecosystem is unmatched for statistical visualization. The learning curve is real, though - budget a few weeks to become productive.
Use raw matplotlib/plotly if you genuinely enjoy programming, need extreme customization, or are building figures into an automated pipeline. You will have full control, and you will earn it through documentation-reading.
Use Plotivy if you know what figure you want but do not want to spend hours on formatting. If "make me a grouped bar chart with error bars, colorblind-safe colors, and Nature formatting" sounds like something you would rather say than code, that is exactly our use case. You describe the figure, Plotivy generates the Python code, and you get a publication-ready export in minutes. The generated code is yours - you can download it, modify it, and run it anywhere.
Chart gallery
PhD-Ready Chart Templates
Browse chart types with publication-quality defaults for your thesis.
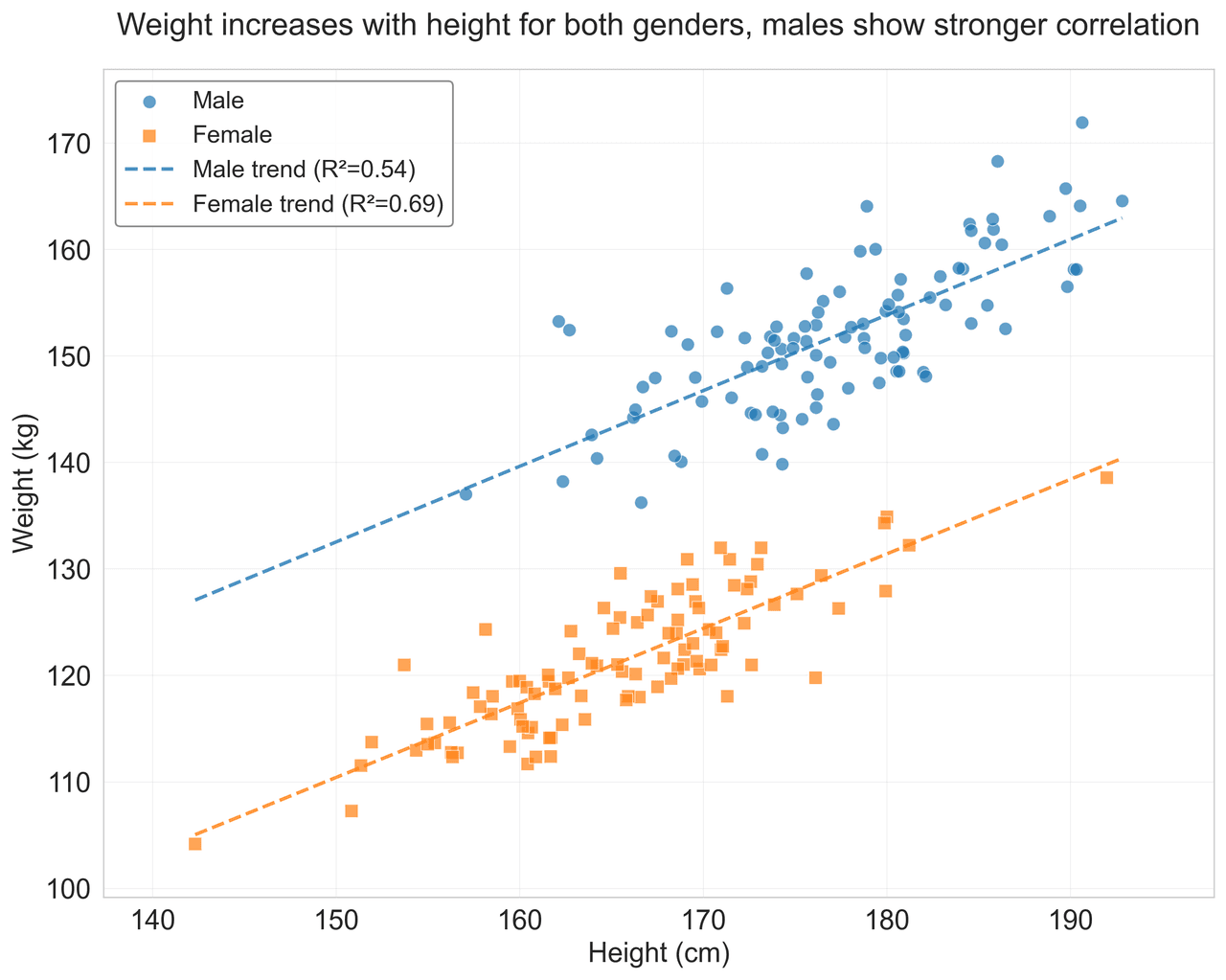
Scatterplot
Displays values for two variables as points on a Cartesian coordinate system.
Sample code / prompt
import matplotlib.pyplot as plt
import numpy as np
from scipy import stats
import pandas as pd
# Generate sample data
np.random.seed(42)
n_samples = 200
height = np.random.normal(170, 8, n_samples)
weight = height * 0.6 + np.random.normal(0, 8, n_samples) - 50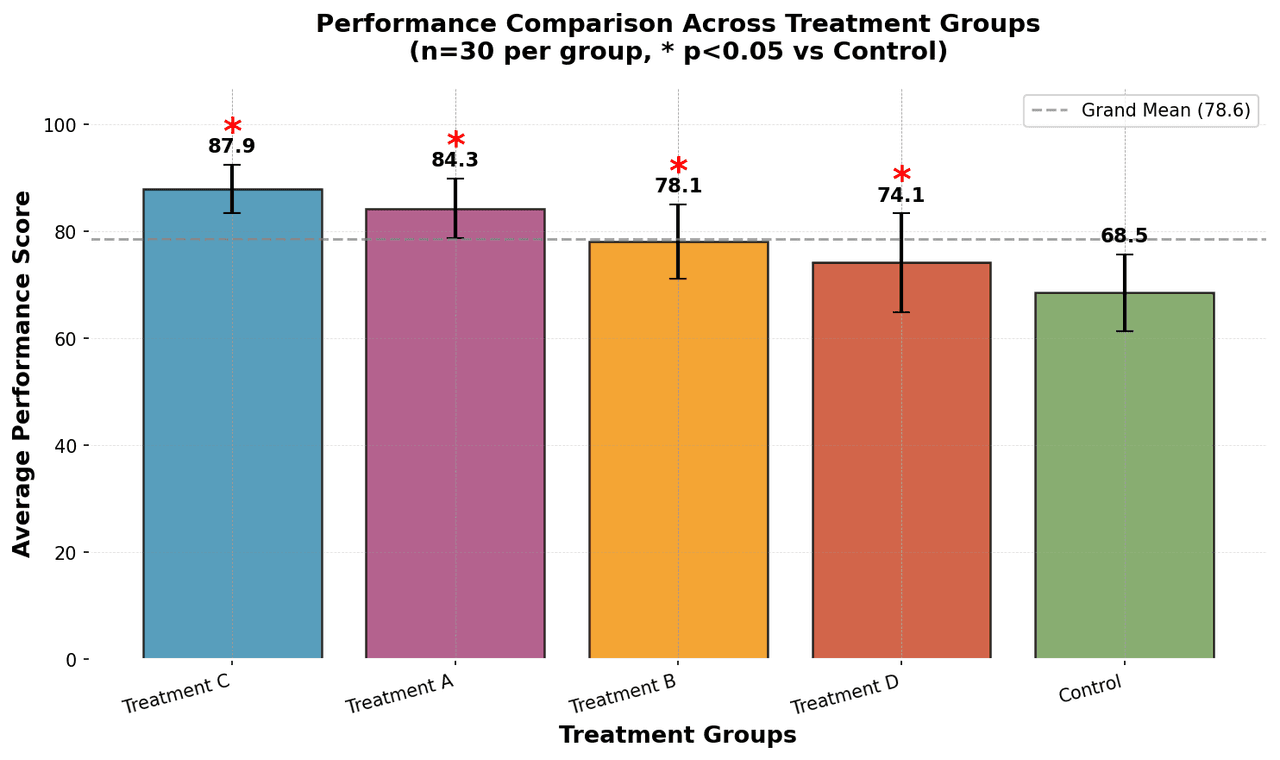
Bar Chart
Compares categorical data using rectangular bars with heights proportional to values.
Sample code / prompt
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
# Generate performance scores for 5 treatment groups
np.random.seed(42)
groups = ['Control', 'Treatment A', 'Treatment B', 'Treatment C', 'Treatment D']
n_samples = 30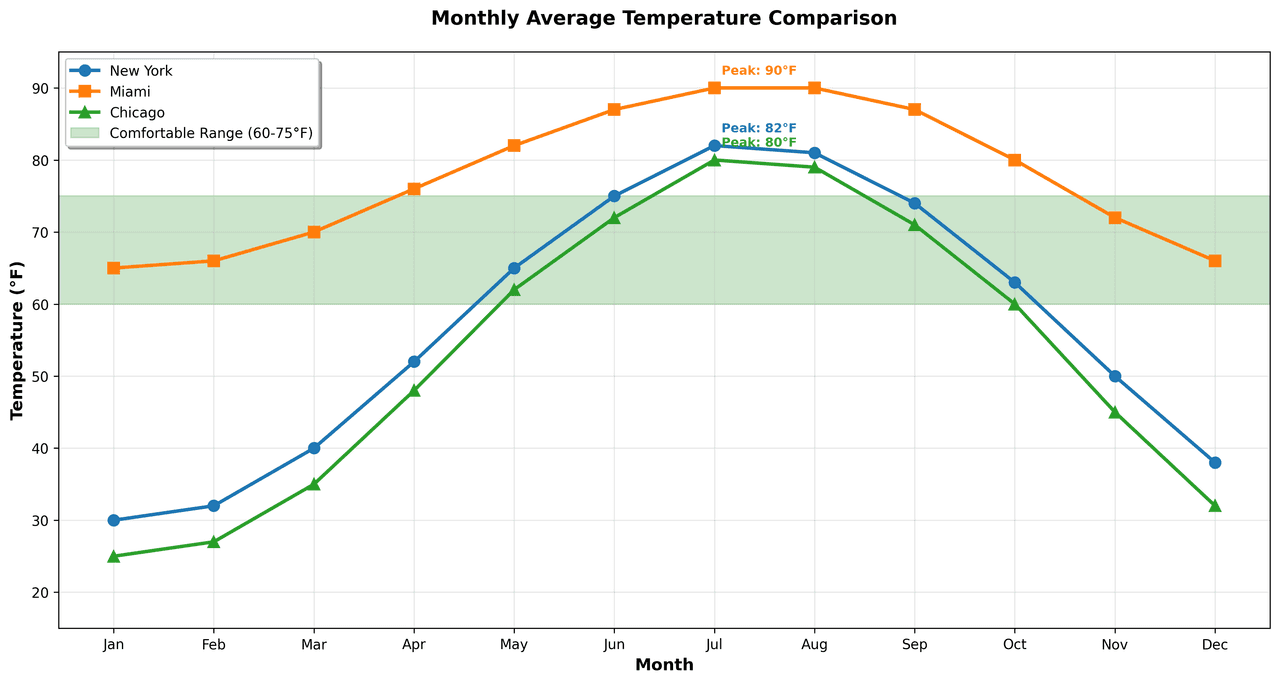
Line Graph
Displays data points connected by straight line segments to show trends over time.
Sample code / prompt
import matplotlib.pyplot as plt
import numpy as np
# Generate temperature data for 3 major US cities over 12 months
months = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun', 'Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec']
nyc = [30, 32, 40, 52, 65, 75, 82, 81, 74, 63, 50, 38]
miami = [65, 66, 70, 76, 82, 87, 90, 90, 87, 80, 72, 66]
chicago = [25, 27, 35, 48, 62, 72, 80, 79, 71, 60, 45, 32]
# Create figure with enhanced styling.png&w=1280&q=70)
Box and Whisker Plot
Displays data distribution using quartiles, median, and outliers in a standardized format.
Sample code / prompt
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
# Generate gene expression data for 4 genotypes
np.random.seed(42)
genotypes = ['WT', 'KO1', 'KO2', 'Mutant']
n_per_group = 20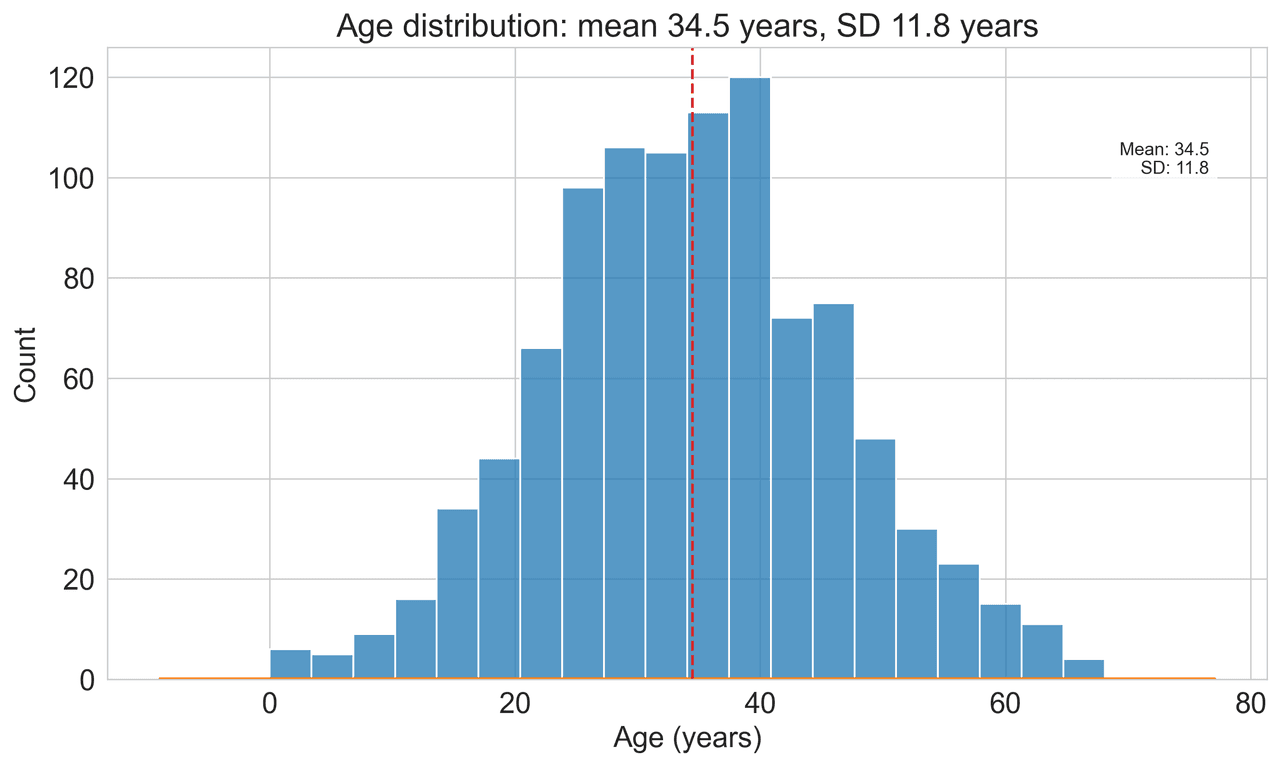
Histogram
Displays the distribution of numerical data by grouping values into bins.
Sample code / prompt
import matplotlib.pyplot as plt
import numpy as np
from scipy.stats import gaussian_kde, skewnorm
# Generate age data with slight right skew
np.random.seed(42)
ages = skewnorm.rvs(a=2, loc=42, scale=15, size=500)
ages = np.clip(ages, 18, 80) # Clip to realistic range
fig, ax = plt.subplots(figsize=(12, 7))Stop Formatting. Start Publishing.
Upload your data and get a publication-ready figure in minutes - not hours. The code is yours, the figure is yours, and your weekend is yours.
Start CreatingYour Figures Should Not Be the Hard Part
You spent months - maybe years - collecting that data. The analysis was the hard part. The experimental design was the hard part. Exporting a plot at the correct DPI with readable fonts and a legend that does not cover your results should not be the thing that keeps you up at 3 AM.
Fix the five disasters above and you will eliminate most of the formatting pain immediately. Build a proper workflow and you will never have another figure emergency before a deadline. And if you want to skip the entire process and go straight from data to publication-ready figure, you know where to find us.
Good luck with the submission. You have got this.
Technique guides scientists read next
scipy.signal.find_peaks guide
Tune prominence and width parameters for robust peak extraction.
Savitzky-Golay smoothing
Reduce noise while preserving peak shape and position.
PCA visualization workflow
Move from high-dimensional measurements to interpretable components.
ANOVA with post-hoc brackets
Add statistically correct pairwise significance annotations.
Found this helpful? Share it with your network.
Experimental Physicist & Photonics Researcher
Hands-on experience in silicon photonics, semiconductor fabrication (DRIE/ICP-RIE), optical simulation, and data-driven analysis. Built Plotivy to help researchers focus on discoveries instead of data struggles.
More about the authorVisualize your own data
Apply the techniques from this article to your own datasets. Upload CSV, Excel, or paste data directly.