Why Your Research Figures Are Not Reproducible - And What To Do About It

Here is an uncomfortable statistic: the majority of figures in published scientific papers cannot be independently recreated from their underlying data. Not because of fraud. Not because researchers are sloppy. But because the pipeline from raw numbers to final PDF is almost never documented well enough for anyone - including the original author six months later - to press a button and get the same output.
This is a quiet, systemic failure in how science communicates results. The paper gets published, the supplemental data gets uploaded to a repository, and the figure itself becomes an orphan - a PNG with no lineage. If you have ever tried to recreate a colleague's figure from their "shared" data folder and failed, you already know the problem.
What You'll Learn
1.The Problem: Figures Without a Paper Trail
2.What Reproducibility Actually Means for Figures
3.Why This Matters More Than Researchers Think
4.The Code-First Approach to Figures
5.Where Plotivy Fits Into This Workflow
6.Five Steps You Can Take This Week
7.The Bar Is Rising - Get Ahead of It
The Problem: Figures Without a Paper Trail
A 2019 study in PLOS Biology attempted to reproduce figures from 50 highly cited papers where the original authors had shared their data. The success rate was dismal. In many cases, the raw data was available, but the exact processing steps - which filters were applied, which outliers were removed, which normalization was used - were either buried in supplemental text or simply missing. The figures were technically "based on" the data but could not be regenerated from it.
This happens for entirely predictable reasons. A researcher opens Excel, manually adjusts axis ranges, picks a colormap by clicking through options, exports at whatever DPI the default suggests, and moves on. The figure looks fine. It passes review. But no record exists of the exact sequence of decisions that created it.
Consider a typical scenario. A postdoc generates a scatter plot in Origin, applies a custom template, manually adds trend lines, and exports a TIFF. Two years later, a reviewer asks for a revised version with different axis labels. The postdoc has left the lab. The Origin project file is on a backup drive no one can find. The figure is effectively frozen in time - unchangeable, unverifiable, unreproducible.
Common ways figure reproducibility breaks down
- GUI-based tools (Excel, Origin, Prism) with no exportable action log
- Manual tweaks: drag-to-resize, click-to-recolor, eyeball axis limits
- Data processed in one tool, plotted in another, with no bridge documented
- Fonts substituted silently across operating systems
- Matplotlib or R version upgrades changing default behaviors
What Reproducibility Actually Means for Figures
Sharing raw data is necessary but nowhere near sufficient. A CSV file on Zenodo does not reproduce a figure. Reproducibility means that someone with your data and your script can run a single command and get a pixel-identical (or semantically identical) output. Every parameter that shaped the visual must be declared explicitly in code.
That list of parameters is longer than most researchers realize:
Parameters that define a figure
figsize - Physical dimensions (inches or cm)
dpi - Resolution (300 for print, 150 for screen)
fontfamily - Arial, Helvetica, or serif fallback
fontsize - Axes labels, tick labels, legend
cmap / palette - Exact colormap with hex values
xlim / ylim - Axis range boundaries
linewidth - Line and marker stroke weight
matplotlib.__version__ - Library version locked
When any of these are left to defaults, you are gambling that the defaults will not change between machines, OS versions, or library updates. Matplotlib 3.7 changed its default color cycle. Plotly 5.x altered tick label formatting. These are silent changes that make your "working" script produce subtly different figures on a different computer.
A reproducible figure script pins every visual decision. It reads raw data from a specified path, applies explicit transformations, and writes a final image file with declared resolution and format. No interactive adjustments. No manual steps. One command, one output.
Updating preview...
Execution Error
Edit the code, then press Run Code to refresh the preview
Preparing preview
Running once automatically on first load
Learn by Experimenting
This is a safe playground for learning! Try changing:
- • Colors: Modify color values to see different palettes
- • Numbers: Adjust sizes, positions, or data ranges
- • Labels: Update titles, axis names, or legends
Edit the code, run it, then open the full data visualization tool to continue with your own dataset.
That script is 18 lines. It encodes every visual decision. A colleague can run it in 2026 and get the same figure you generated in 2025, provided the pinned library versions are installed. That is reproducibility.
Why This Matters More Than Researchers Think
Figure reproducibility is not an abstract ideal pursued by open-science advocates. It has immediate, practical consequences that hit researchers at the worst possible moments.
Reviewer requests. The average paper goes through at least one round of revisions, and reviewers routinely ask for figure modifications: "Please use a log scale on the y-axis," "Combine panels A and B into a single figure," "Add error bars." If the figure was made in a GUI with no saved workflow, each modification means starting from scratch, manually re-applying every aesthetic choice. If it was made with a script, you change one line and re-run.
Journal mandates. Journals are tightening requirements. Nature now encourages source data files alongside figures. eLife requires figures to be "computationally reproducible" where feasible. PLOS journals have mandatory data availability statements. The trajectory is clear: within five years, submitting the script that generates your figures will likely be as standard as submitting supplemental data is now.
Try it
Try it now: turn this method into your next figure
Apply the same approach to your own dataset and generate clean, publication-ready code and plots in minutes.
Open in Plotivy Analyze →Newsletter
Get a weekly Python plotting tip
One concise tip each week for cleaner, faster scientific figures. Built for researchers who publish.
Your future self. Every researcher who has returned to a project after six months knows the feeling: staring at a figure in a draft, unable to remember how it was made. Which version of the data? Which normalization? Was the outlier at row 47 included or excluded? Without a script, the answers live only in memory, and memory is unreliable.
Lab continuity. Graduate students leave. Postdocs move on. If the only person who knows how Figure 3 was generated has graduated, the lab has a problem. A well-documented figure script in a shared repository is institutional knowledge that survives personnel changes.
The Code-First Approach to Figures
The fix is not complicated, but it requires a shift in habit. The principle is simple: every figure in a paper should have a script that generates it from raw data in a single command. No intermediate manual steps. No "open the file in Excel first." Raw data in, final figure out.
This is what software engineers call a build pipeline, and it applies directly to research figures. Your project directory should look something like this:
Updating preview...
Execution Error
Edit the code, then press Run Code to refresh the preview
Preparing preview
Running once automatically on first load
Learn by Experimenting
This is a safe playground for learning! Try changing:
- • Colors: Modify color values to see different palettes
- • Numbers: Adjust sizes, positions, or data ranges
- • Labels: Update titles, axis names, or legends
Edit the code, run it, then open the full data visualization tool to continue with your own dataset.
The Makefile (or a simple shell script) ties it together:
Updating preview...
Execution Error
Edit the code, then press Run Code to refresh the preview
Preparing preview
Running once automatically on first load
Learn by Experimenting
This is a safe playground for learning! Try changing:
- • Colors: Modify color values to see different palettes
- • Numbers: Adjust sizes, positions, or data ranges
- • Labels: Update titles, axis names, or legends
Edit the code, run it, then open the full data visualization tool to continue with your own dataset.
Run make all and every figure regenerates. Change the data, re-run, and the figures update. Check the entire directory into Git, and you have a version-controlled record of every figure at every stage of the project. A reviewer asks for a change? Edit the script, commit, re-run. The diff shows exactly what changed and why.
Version control for figures is profoundly underused in academia. In industry, no one would ship a product without version-controlled build artifacts. Yet researchers routinely publish papers where the figures exist as standalone files with names like fig2_final_FINAL_v3_revised.png. Git solves this entirely.
Where Plotivy Fits Into This Workflow
If the code-first approach is the goal, the obvious barrier is that writing figure scripts from scratch is slow. Most researchers are not software developers. They know enough Python or R to be functional, but writing airtight, publication-quality bar charts, heatmaps, or multi-panel layouts from memory is not where their time should go.
This is the specific problem Plotivy was designed to solve. When you generate a figure in Plotivy, the platform produces two outputs simultaneously: the rendered figure and the complete Python script that created it. The script is not an afterthought or an export option buried in a menu. It is the primary artifact. The figure is just what the script produces.
That distinction matters. It means every figure you create through Plotivy is reproducible by default. You can download the script, drop it into your project's scripts/ directory, pin the library versions, and commit it alongside your data. Six months from now - or six years - anyone can re-run it.
The AI-assisted workflow also handles the tedious parts: choosing appropriate error bar styles, setting journal-compliant font sizes, applying colorblind-safe palettes. You describe what you need in plain language, and the generated script encodes all of those decisions explicitly. No hidden defaults, no GUI state that vanishes when you close the window.
Five Steps You Can Take This Week
You do not need to overhaul your entire workflow at once. These five actions are concrete, incremental, and each one independently improves your figure reproducibility.
- Create a
figures/andscripts/directory in your current project. Separate figure-generating code from analysis notebooks. Each script should produce exactly one figure and be named to match:fig1_scatter.pyproducesfig1_scatter.png. - Add a
requirements.txtwith pinned versions. Runpip freezeand save the output. At minimum, pin matplotlib, numpy, and pandas. This single file prevents 90% of "it worked on my machine" failures. - Replace one GUI-made figure with a script. Pick the simplest figure in your current draft - a basic line chart or scatter plot. Rewrite it as a Python script with explicit parameters. Use Plotivy to generate the initial code if writing it manually feels slow.
- Put your project under version control. If you are not already using Git, initialize a repository today. Commit raw data, scripts, and generated figures. Every figure revision becomes a trackable commit, not a renamed file.
- Write a one-line regeneration command. Whether it is a Makefile, a shell script, or a README instruction, document how to regenerate all figures from raw data. The standard: a new lab member should be able to clone the repository and reproduce every figure within 10 minutes.
None of these steps require advanced programming skill. None require changing your analysis methodology. They are organizational habits that compound over time. A project that follows these conventions from the start will save dozens of hours during revisions - and will still be fully reproducible when the paper is cited a decade from now.
Chart gallery
Reproducible Chart Templates
Browse chart types with full Python source code for reproducible workflows.
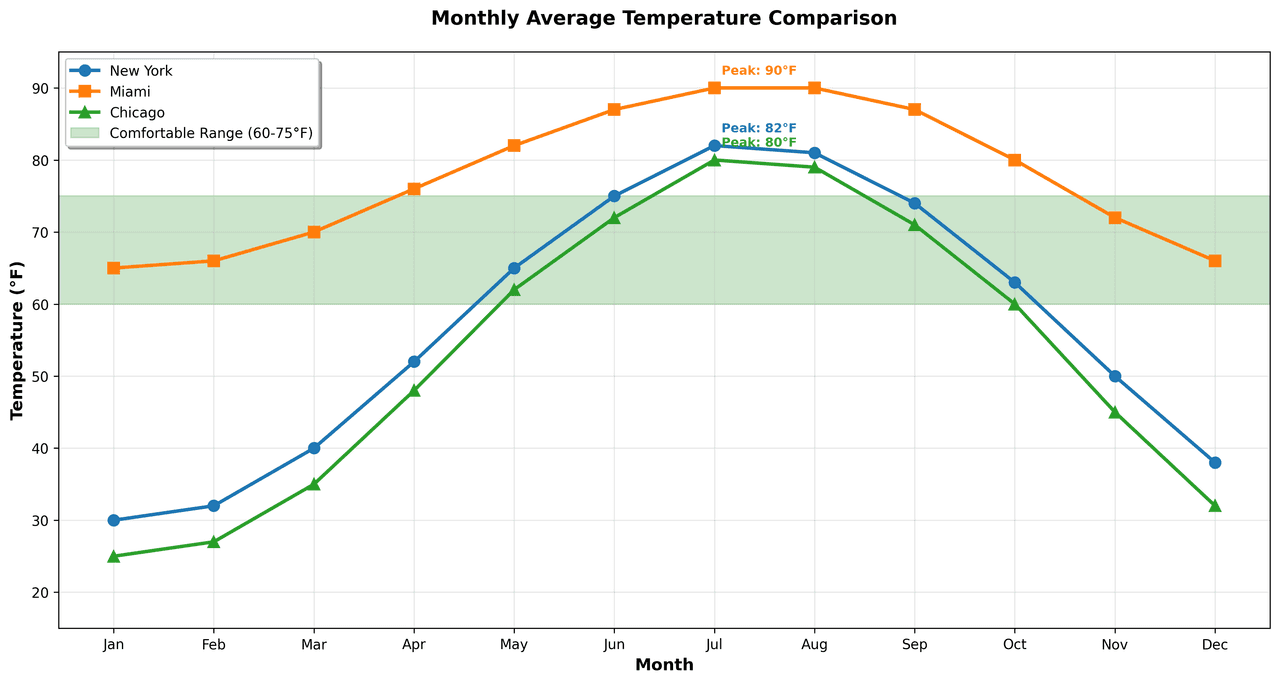
Line Graph
Displays data points connected by straight line segments to show trends over time.
Sample code / prompt
import matplotlib.pyplot as plt
import numpy as np
# Generate temperature data for 3 major US cities over 12 months
months = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun', 'Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec']
nyc = [30, 32, 40, 52, 65, 75, 82, 81, 74, 63, 50, 38]
miami = [65, 66, 70, 76, 82, 87, 90, 90, 87, 80, 72, 66]
chicago = [25, 27, 35, 48, 62, 72, 80, 79, 71, 60, 45, 32]
# Create figure with enhanced styling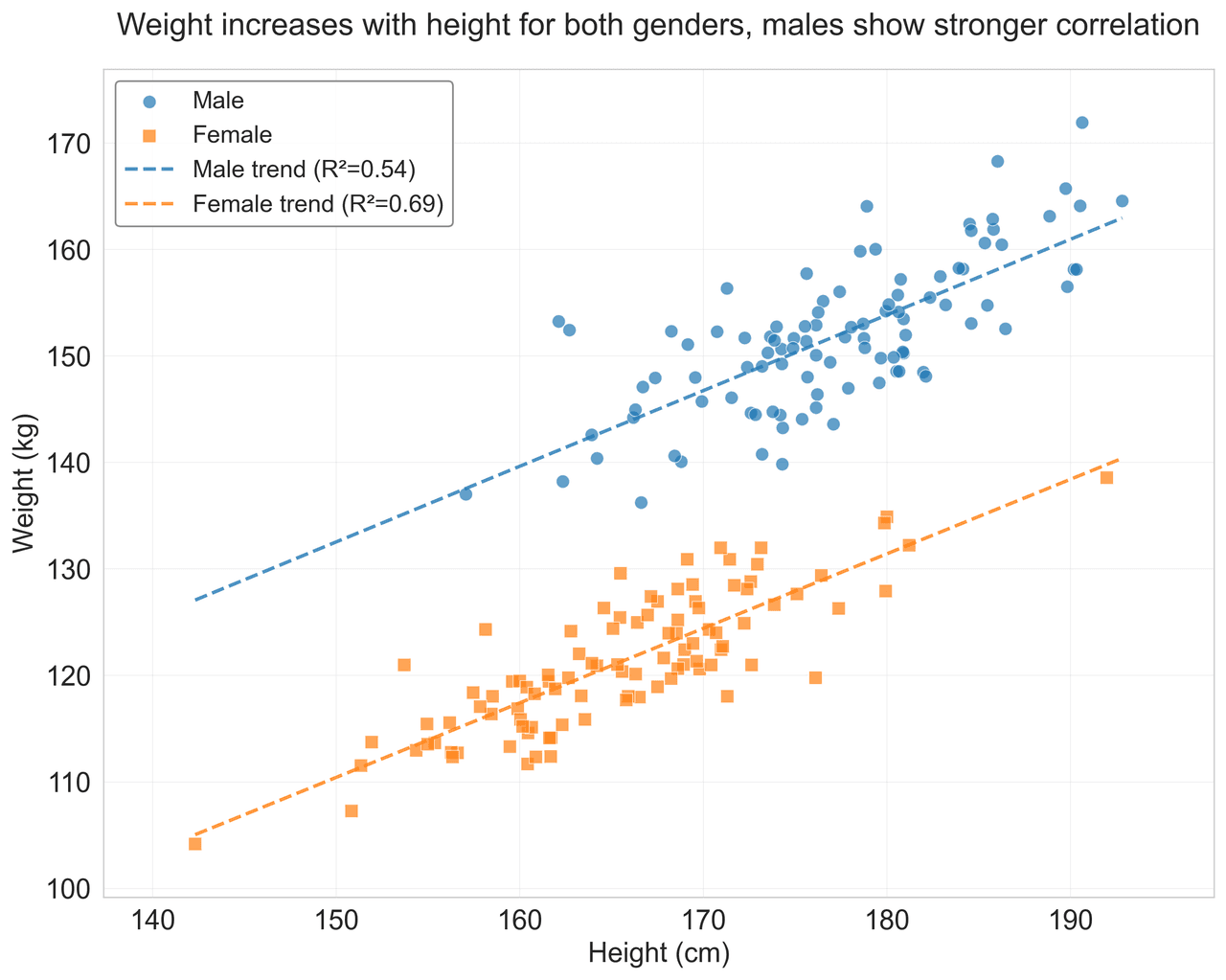
Scatterplot
Displays values for two variables as points on a Cartesian coordinate system.
Sample code / prompt
import matplotlib.pyplot as plt
import numpy as np
from scipy import stats
import pandas as pd
# Generate sample data
np.random.seed(42)
n_samples = 200
height = np.random.normal(170, 8, n_samples)
weight = height * 0.6 + np.random.normal(0, 8, n_samples) - 50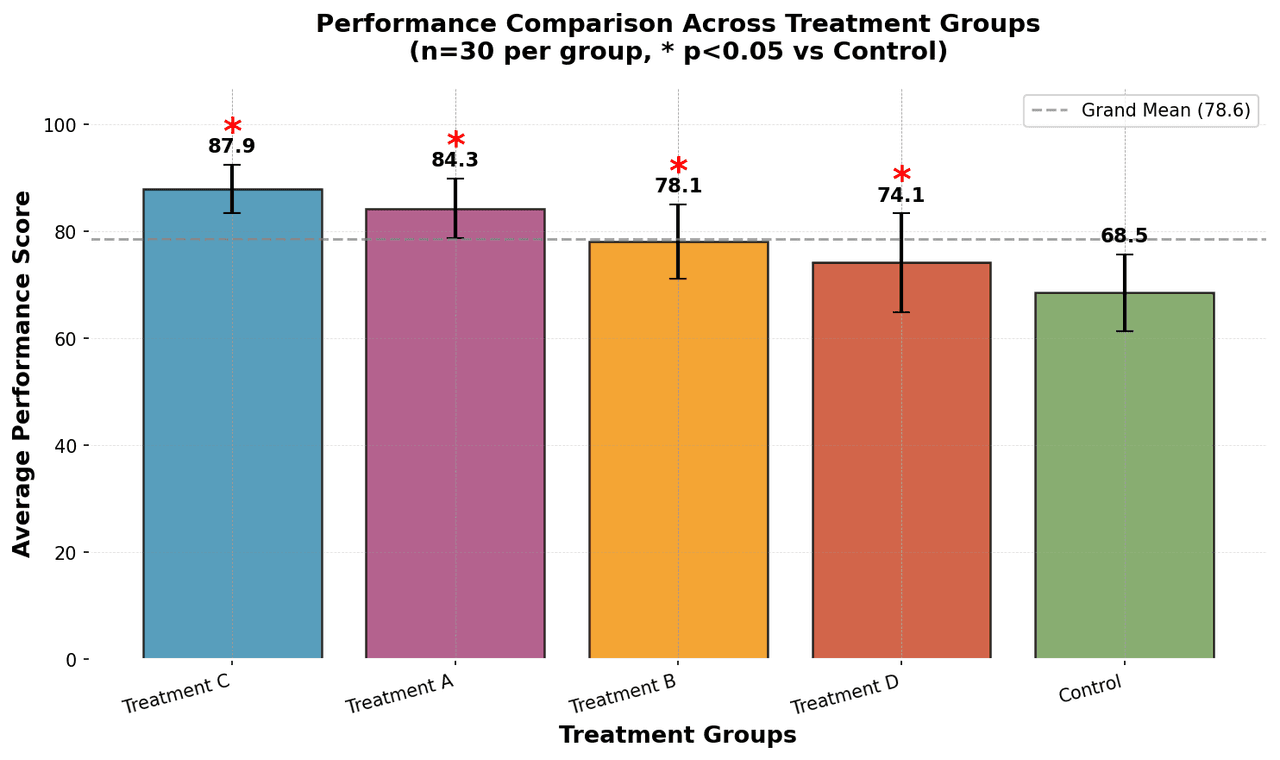
Bar Chart
Compares categorical data using rectangular bars with heights proportional to values.
Sample code / prompt
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
# Generate performance scores for 5 treatment groups
np.random.seed(42)
groups = ['Control', 'Treatment A', 'Treatment B', 'Treatment C', 'Treatment D']
n_samples = 30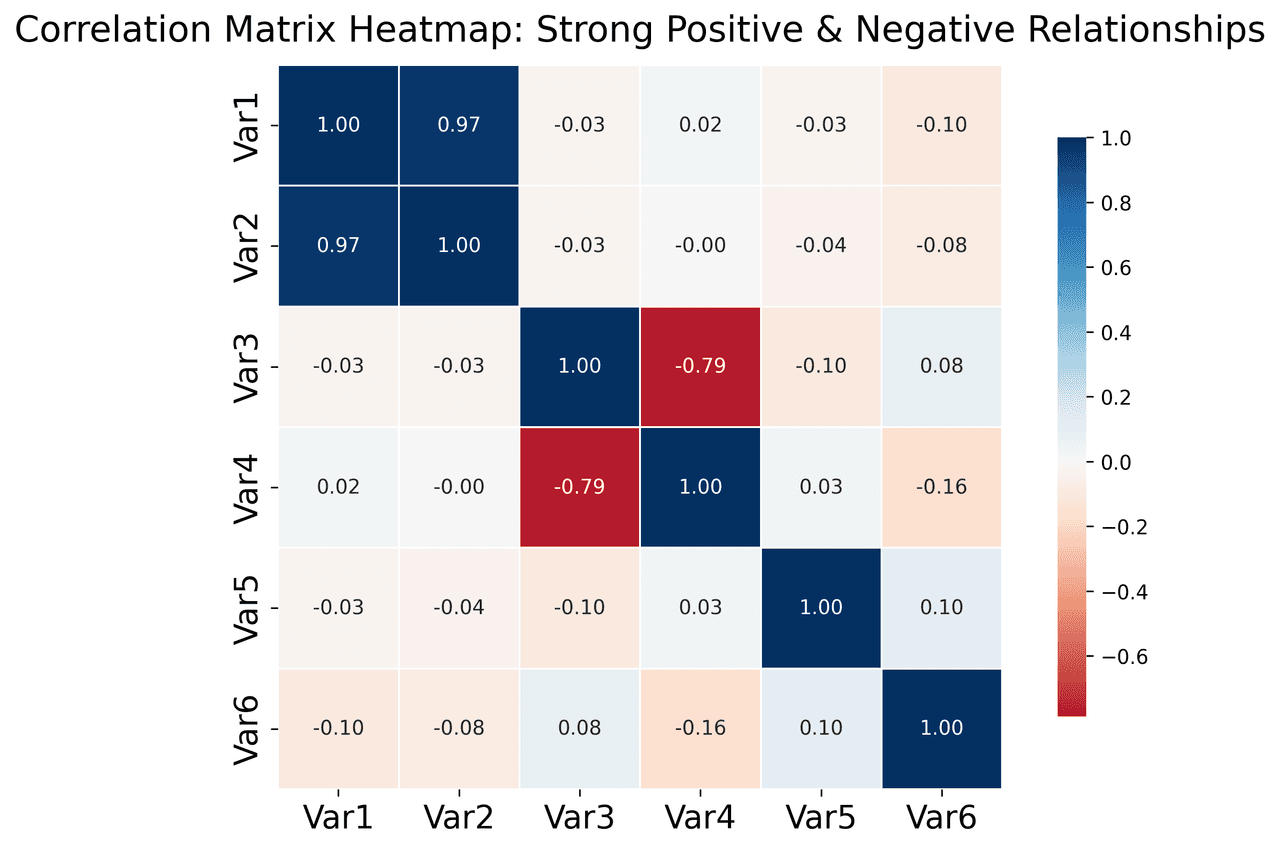
Heatmap
Represents data values as colors in a two-dimensional matrix format.
Sample code / prompt
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
# Create correlation matrix for financial metrics
metrics = ['Revenue', 'Profit', 'Expenses', 'ROI', 'Customers', 'AOV', 'Marketing', 'Employees']
correlation_data = np.array([
[1.00, 0.85, -0.45, 0.72, 0.88, 0.65, 0.72, 0.55],
[0.85, 1.00, -0.78, 0.92, 0.75, 0.58, 0.63, 0.48],.png&w=1280&q=70)
Box and Whisker Plot
Displays data distribution using quartiles, median, and outliers in a standardized format.
Sample code / prompt
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
# Generate gene expression data for 4 genotypes
np.random.seed(42)
genotypes = ['WT', 'KO1', 'KO2', 'Mutant']
n_per_group = 20Start Making Reproducible Figures Now
Upload your data and get a publication-ready plot with the full Python script - ready to commit to your repository.
Start CreatingThe Bar Is Rising - Get Ahead of It
Reproducibility is moving from "nice to have" to "required." Funders are mandating open data. Journals are mandating source code. Preprint servers are adding reproducibility badges. The researchers who build reproducible habits now will not scramble to comply later - they will already be there.
The irony is that reproducible figures are also easier to work with. They are faster to revise. They survive personnel changes. They make collaboration straightforward instead of painful. The upfront cost of writing a script instead of clicking through a GUI pays for itself the first time a reviewer asks for a change.
Your figures are the most visible part of your research. They are what gets shared on social media, what appears in presentations, and what readers remember. Making them reproducible is not just good practice - it is respect for the work that produced them.
Technique guides scientists read next
scipy.signal.find_peaks guide
Tune prominence and width parameters for robust peak extraction.
Savitzky-Golay smoothing
Reduce noise while preserving peak shape and position.
PCA visualization workflow
Move from high-dimensional measurements to interpretable components.
ANOVA with post-hoc brackets
Add statistically correct pairwise significance annotations.
Found this helpful? Share it with your network.
Experimental Physicist & Photonics Researcher
Hands-on experience in silicon photonics, semiconductor fabrication (DRIE/ICP-RIE), optical simulation, and data-driven analysis. Built Plotivy to help researchers focus on discoveries instead of data struggles.
More about the authorVisualize your own data
Apply the techniques from this article to your own datasets. Upload CSV, Excel, or paste data directly.